Anand Singh Rathore
I'm Designer
I am a doctoral researcher at the Indraprastha Institute of Information Technology Delhi (IIIT-Delhi), working under the supervision of Prof. Gajendra P.S. Raghava. I hold a master’s degree in Zoology (2020–2022) and have since transitioned into the field of Data Science, driven by the transformative potential of Machine Learning and Artificial Intelligence. My current research focuses on peptide-based therapeutics, particularly exploring their druggability using computational approaches. Computational Biology, being a multidisciplinary field, allows for innovation at the intersection of biology, data science, and artificial intelligence. This fusion offers vast research opportunities to address complex biological challenges and advance therapeutic development.

Zoologist Turned Computational Biologist
Exploring the crossroads of biology and computation to unlock new possibilities in precision medicine and therapeutic design.
- ORCID: 0009-0004-8907-7174
- DOB: 2 July 1998
- Phone: +91 8819919830
- City: New Delhi, India
- Google Scolar: Anand Singh Rathore
- Degree: M.Sc. Zoology
- Email 1: anandr@iiitd.ac.in
- Email 2: anandrathoreindia@gmail.com
I actively work on building and deploying machine learning pipelines for bioinformatics applications such as toxicity prediction, motif analysis, and peptide screening. My toolkit spans Python, R, shell scripting, and technologies like BLAST, IEDB, SwissProt, and ESM-based models. I’m also exploring the frontiers of Quantum Machine Learning and Explainable AI to make bioinformatics models more transparent and insightful. My journey is driven by curiosity, interdisciplinary collaboration, and a desire to make meaningful contributions to biomedical research.
Technical Expertise
A curated showcase of my core competencies in computational biology and data science
Programming & Development
AI & Machine Learning
Bioinformatics Tools
Resume
Research scholar in Computational Biology with strong academic background, practical laboratory training, and diverse experience in machine learning, proteiomics, genomics, and bioinformatics.
Education
Ph.D. in Computational Biology (Pursuing)
2022 – PresentIndraprastha Institute of Information Technology (IIIT-Delhi), India
Working under the supervision of Prof. (Dr) GPS Raghava on machine learning applications in peptide-based therapeutics.
M.Sc. in Zoology
2020 – 2022Hansraj College, University of Delhi, Delhi, India
CGPA: 7.3
Acquired in-depth knowledge in animal physiology, animal endocrinology, molecular biology, developmental biology, and immunology. Gained hands-on experience with biological techniques such as microscopy, gel electrophoresis, and basic bioinformatics tools.
B.Sc. in Life Sciences
2017 – 2020Sriyut College, APS University, Rewa, Madhyapradesh, India
Percentage: 72.33%
Built a strong foundation in cell biology, genetics, biochemistry, and microbiology. Developed analytical and laboratory skills, and was introduced to research methodologies and scientific writing.
Achievements
CSIR-UGC NET JRF
June 2022All India Rank: 79 (Percentile: 99.67)
DBT-JRF
2022Qualified under Category-I
GATE XL
2021All India Rank: 409 | Score: 598
GATE BT
2022All India Rank: 906 | Score: 454
Research & Lab Experience
Hands-on Laboratory Training
Jul – Sep 2021Prof. Neeta Sehgal's Lab, Dept. of Zoology, University of Delhi
- Agarose Gel Electrophoresis, SDS-PAGE, PCR, RNA Isolation
- Western Blotting, Real-Time PCR, DNA & Protein Purification
Antimicrobial Peptide & Database Learning
Sep – Dec 2021Under Asst. Prof. Vikash Sood, Jamia Hamdard
- Peptide data extraction, Venn diagrams, Heatmaps, Clustering
In Silico Biology Virtual Internship
Sep 2021Under Dr. Shobhana Manoharan, Biosrishti
- BLAST, FASTA, Phylogenetics, Protein Docking (PatchDock, AutoDock)
- Structure Prediction (Phyre2, SOPMA), Visualization (Chimera, RasMol)
Workshops & Conferences
IEDB Virtual User Workshop
La Jolla Institute for Immunology, USA
Python for Biology Workshop
INSCR 2021
International Conferences
Participated in conferences focusing on Cancer Biology, Nanomedicine, and Genomics
Certifications
(Meta) Genomics & Bioinformatics
Phixgen Pvt. Ltd.
Immunological Techniques
BCAS, University of Delhi
Secured First Merit Position
Publications
Peer-reviewed research articles, preprints, and scientific contributions showcasing our work in computational biology and bioinformatics.
Published Articles
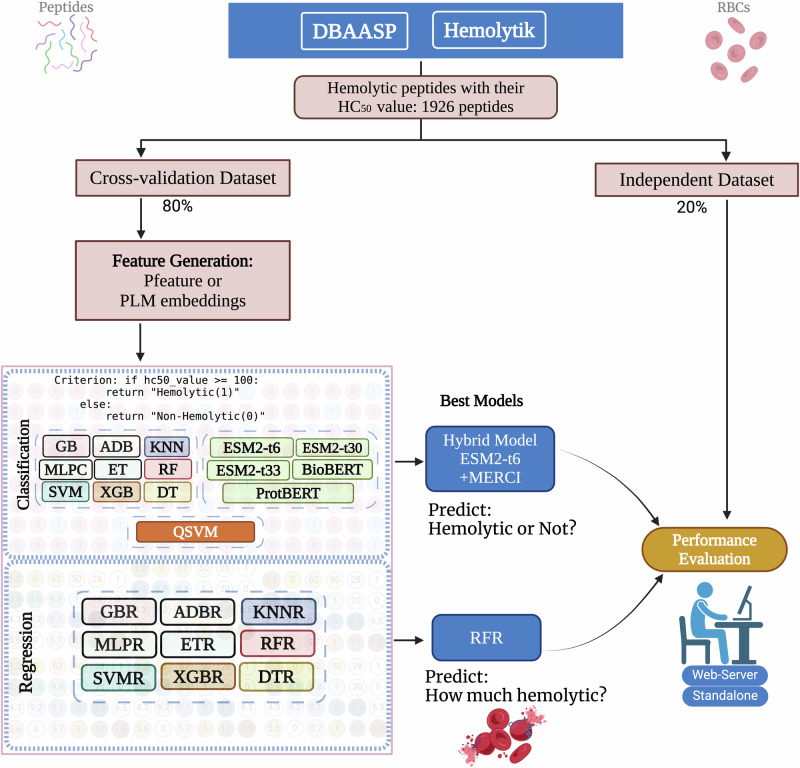
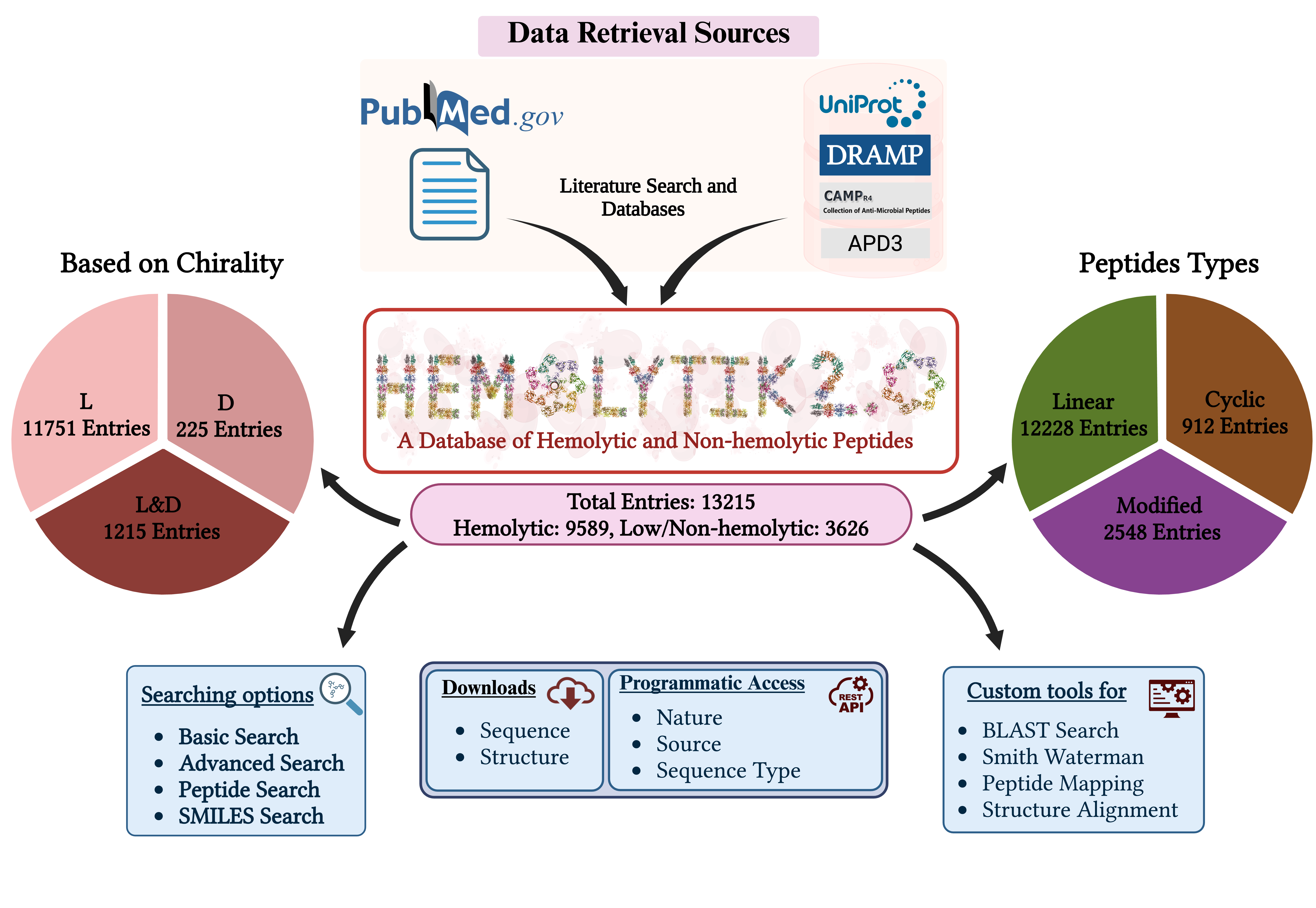
Hemolytik2: An Updated Database of Hemolytic and Non-hemolytic Peptides
Prediction of hemolytic peptides using machine learning approaches
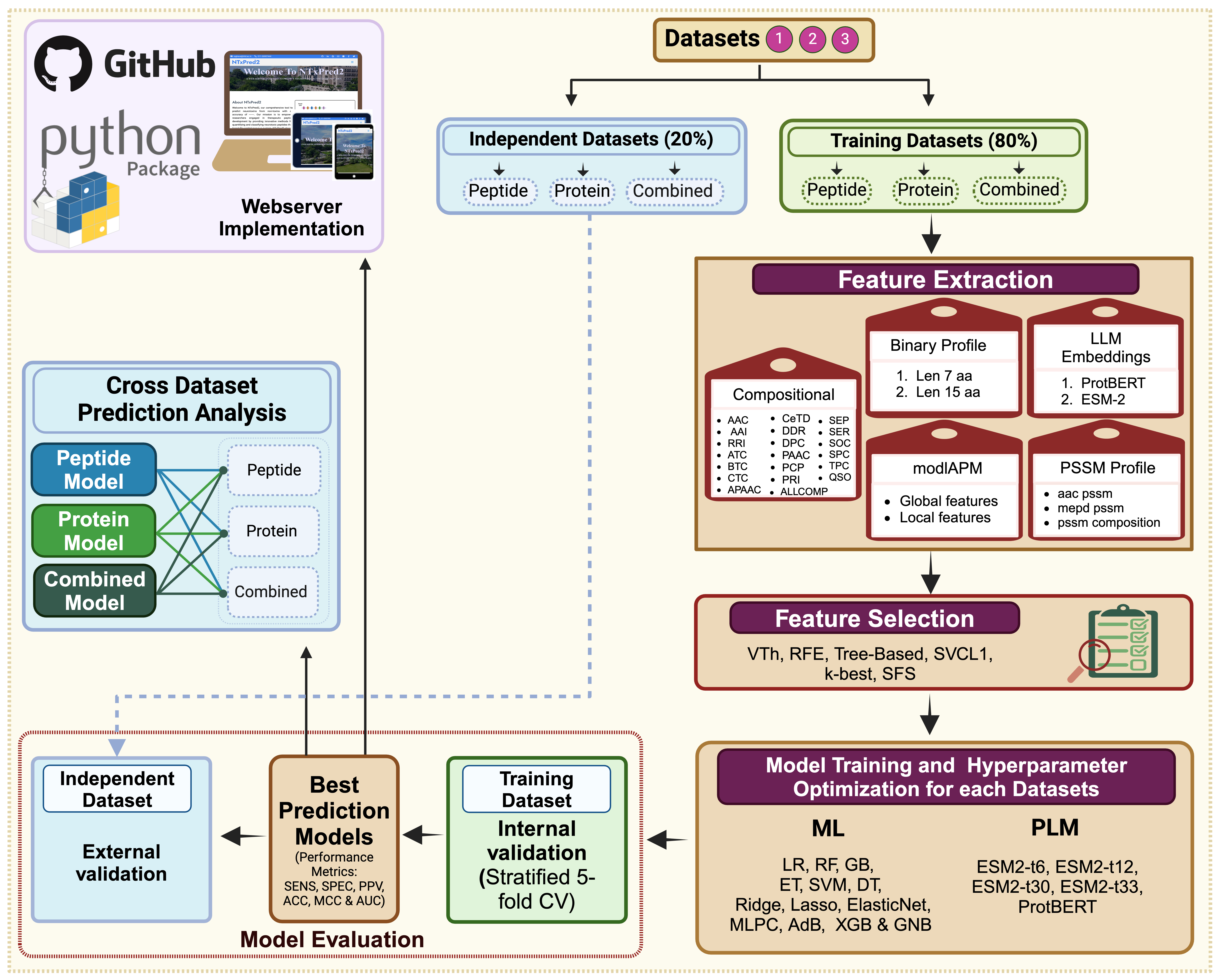
NTxPred2: A large language model for predicting neurotoxins
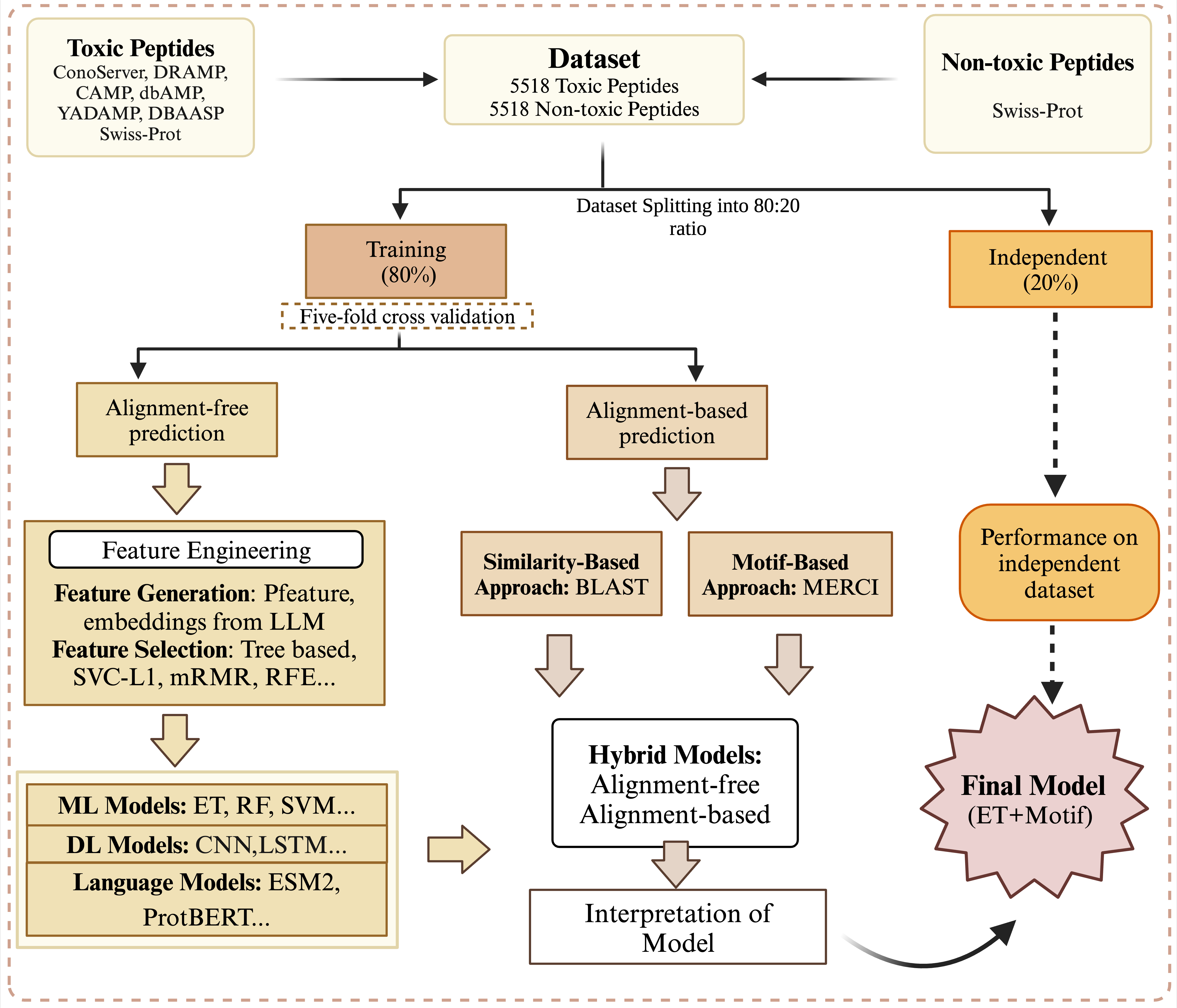
ToxinPred 3.0: An improved method for predicting toxicity of peptides and proteins
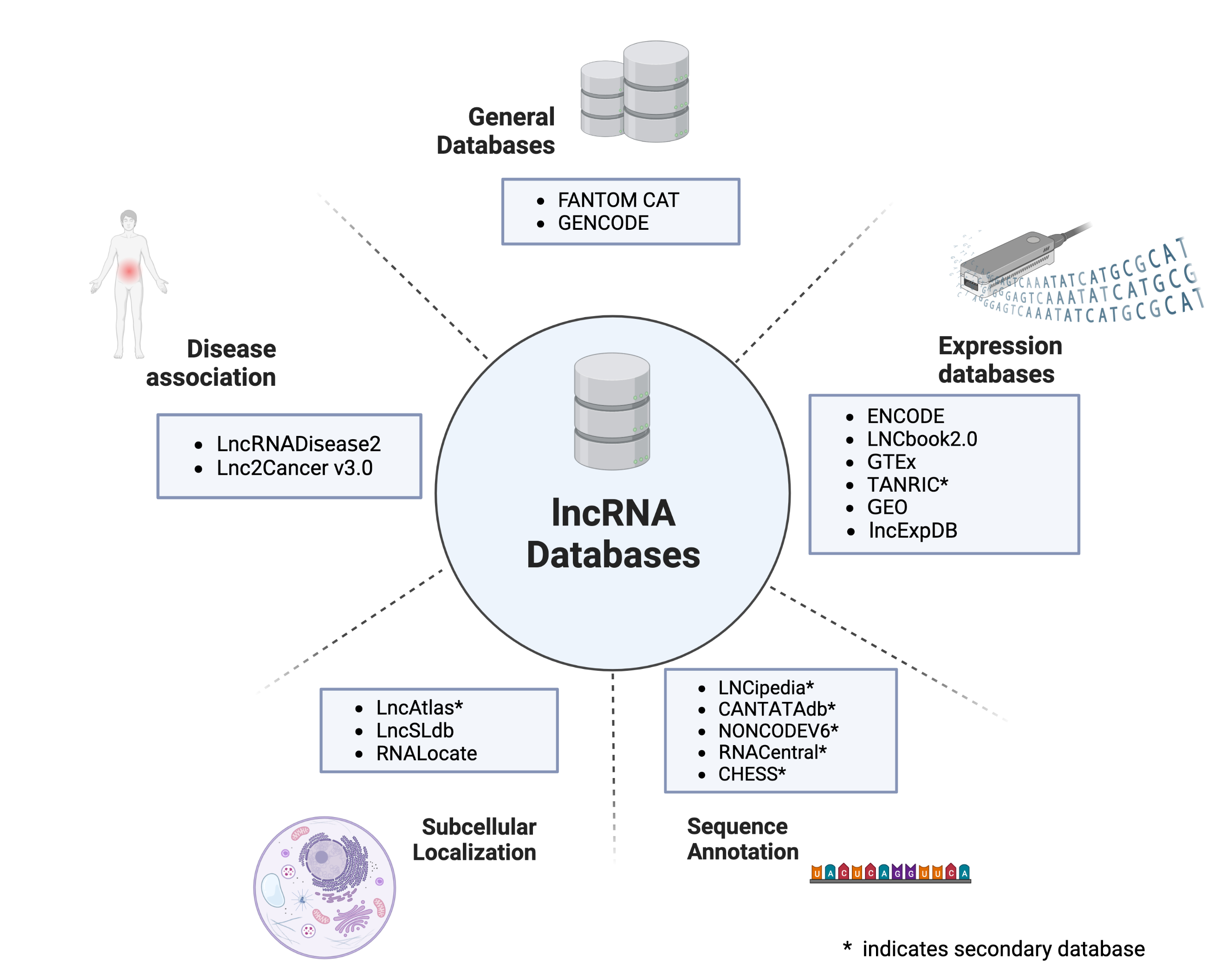
Compilation of resources on subcellular localization of RNA
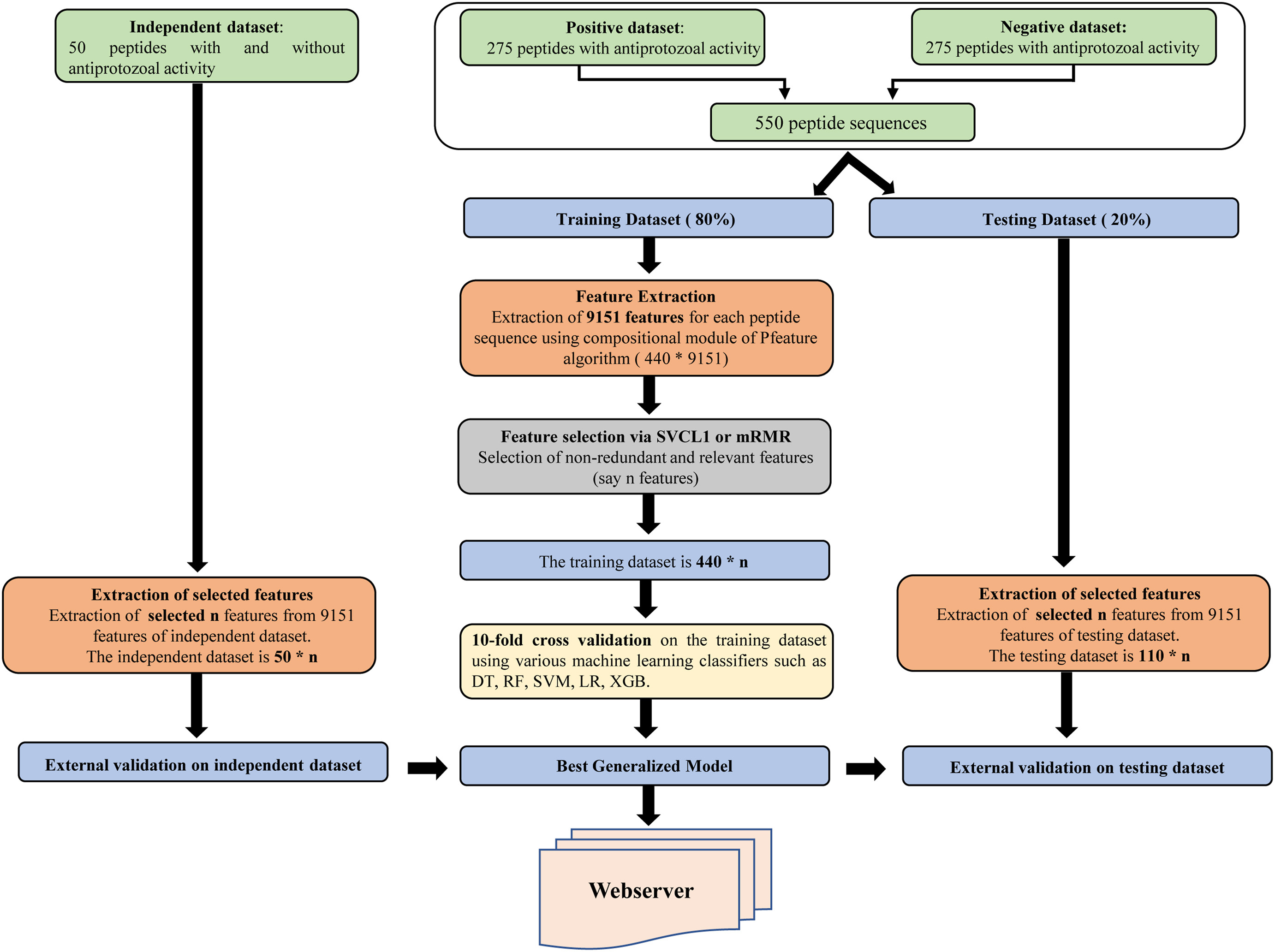
Antiprotozoal peptide prediction using machine learning approaches
Preprints
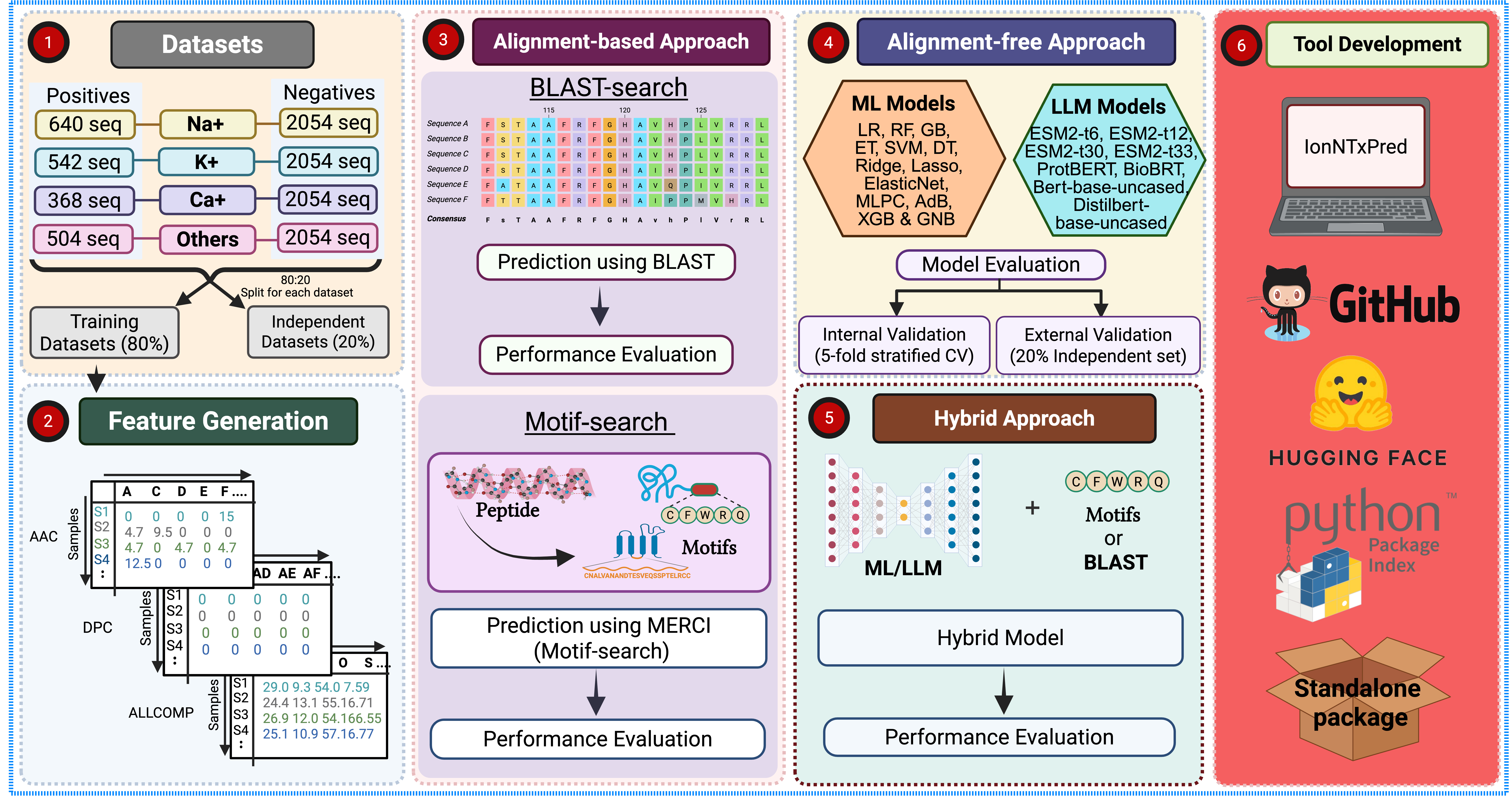
A PLM-Based Method for Predicting Protein Ion Channel Modulators for Drug Discovery and Safety Evaluation
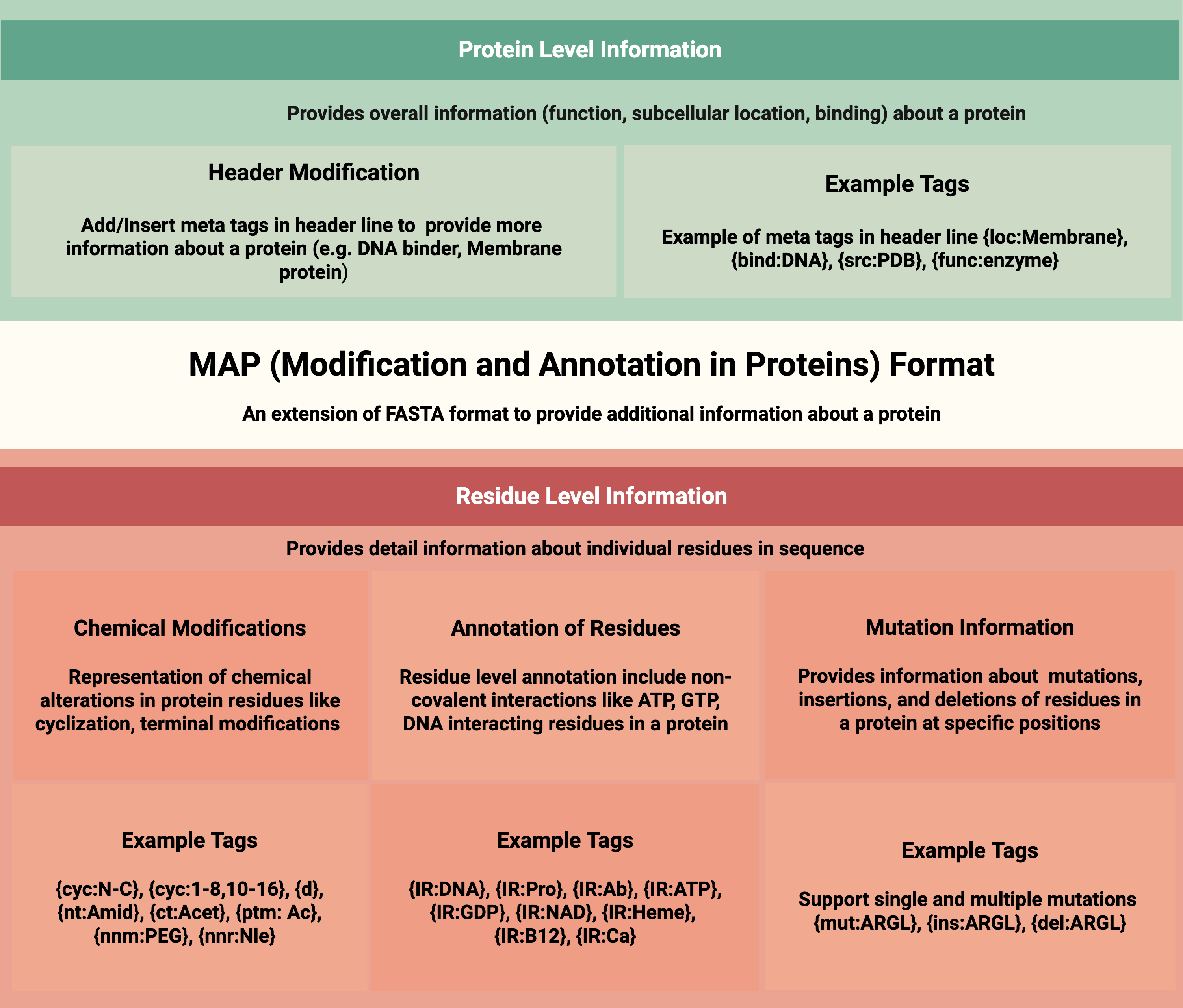
MAP format for representing chemical structures as text for large language models
Webservers
This section showcases a collection of web-based resources and tools developed to facilitate peptide-based therapeutic research. These include predictive models, curated databases, and computational methods for analyzing bioactive peptides.
- All
- Tool
- Database
- Method
- Review
- Book Chapter
Research Areas
My research spans a variety of domains in bioinformatics, computational biology, and artificial intelligence to address challenges in health and life sciences.
Machine Learning in Bioinformatics
Development of predictive models for biological data using supervised and unsupervised learning techniques.
Peptide Therapeutics
Design and prediction of therapeutic peptides for antimicrobial, anticancer, and hemolytic activities.
Toxin and Allergen Prediction
Computational tools for the identification of toxic and allergenic proteins and peptides.
Vaccine Informatics
Epitope mapping and computational vaccine design using immunoinformatics approaches.
AI for Precision Medicine
Leveraging artificial intelligence for personalized diagnosis and treatment strategies.
Web Server and Tool Development
Creating user-friendly, freely available web interfaces and software tools for the scientific community's access to predictive models.
Genomics and Sequence Analysis
Exploration of genomic datasets to uncover gene functions, mutations, and their role in disease and evolution.
My Contributions
Impact-driven work spanning knowledge creation, mentorship, community building, and societal advancement through computational biology.
Generation of Knowledge
Developed advanced machine learning and transformer-based models for peptide toxicity prediction, enabling safer therapeutic design.
Mentorship & Teaching
Guided students in biomedical machine learning courses and research methodology, fostering analytical skills and collaborative spirit.
Community Building
Promoted open science through resource sharing, interdisciplinary collaboration, and active participation in scientific discourse.
Societal Impact
Created accessible computational tools for safer peptide therapeutics, advancing public health objectives through open science.
Cumulative Impact
Journey in Focus
Capturing milestones, collaborations, and memorable moments from my academic and research path.
Contact
Feel free to reach out with any questions or collaboration ideas.
Address
A318 R & D Building, IIIT-Delhi
Call
+91 8819919830
anandr@iiitd.ac.in